- Methodology article
- Open access
- Published:
Genomic profiling of plastid DNA variation in the Mediterranean olive tree
BMC Plant Biology volume 11, Article number: 80 (2011)
Abstract
Background
Characterisation of plastid genome (or cpDNA) polymorphisms is commonly used for phylogeographic, population genetic and forensic analyses in plants, but detecting cpDNA variation is sometimes challenging, limiting the applications of such an approach. In the present study, we screened cpDNA polymorphism in the olive tree (Olea europaea L.) by sequencing the complete plastid genome of trees with a distinct cpDNA lineage. Our objective was to develop new markers for a rapid genomic profiling (by Multiplex PCRs) of cpDNA haplotypes in the Mediterranean olive tree.
Results
Eight complete cpDNA genomes of Olea were sequenced de novo. The nucleotide divergence between olive cpDNA lineages was low and not exceeding 0.07%. Based on these sequences, markers were developed for studying two single nucleotide substitutions and length polymorphism of 62 regions (with variable microsatellite motifs or other indels). They were then used to genotype the cpDNA variation in cultivated and wild Mediterranean olive trees (315 individuals). Forty polymorphic loci were detected on this sample, allowing the distinction of 22 haplotypes belonging to the three Mediterranean cpDNA lineages known as E1, E2 and E3. The discriminating power of cpDNA variation was particularly low for the cultivated olive tree with one predominating haplotype, but more diversity was detected in wild populations.
Conclusions
We propose a method for a rapid characterisation of the Mediterranean olive germplasm. The low variation in the cultivated olive tree indicated that the utility of cpDNA variation for forensic analyses is limited to rare haplotypes. In contrast, the high cpDNA variation in wild populations demonstrated that our markers may be useful for phylogeographic and populations genetic studies in O. europaea.
Background
In the last decades, major technical innovations have allowed a rapid development of various methods for genomic analysis. These have led to applications ranging from phylogeographical reconstructions to forensic analyses and species identification [1, 2]. In plants, many studies have focused on the organelle genomes (i.e., plastid DNA - cpDNA - and mitochondrial DNA - mtDNA) for six major reasons: (i) these genomes are usually uniparentally inherited (either from the mother or the father) and thus allow for investigations of gene dispersal by seeds or pollen without recombination effect [3]; (ii) their haploid nature facilitates their sequencing and usually does not require cloning; (iii) such genomes are more prone to stochastic events because their effective population size is half that of diploid genomes, allowing a more accurate detection of evolutionary events such as a long persistence of relict populations in refuge zones during last glaciations [4]. In addition the dispersion of maternally inherited genomes (due to the seed dissemination only) occurs at shorter geographic distances than for nuclear genomes. The consequence of a reduced gene dispersal and high genetic drift in organelle genomes is a generally pronounced geographic structure, which facilitates phylogeographic analyses as well as tracing the origins of cultivated species or invasive populations [3]; (iv) they exhibit a high number of identical copies per cell [5], which may represent a significant advantage for forensic analyses; (v) they are circular and protected by a double-membrane envelope, which makes them resistant to exonucleases and less prone to endonuclease degradation (another advantage for forensics; [6]); and (vi) they exhibit a lower mutation rate than nuclear genomes [7, 8], and such stability is generally required for traceability analyses (although see below).
The olive tree (Olea europaea, Oleaceae) is among the oldest woody crops, and nowadays represents one of the major cultivated species in the Mediterranean area [9]. The origins of this species have been recently investigated using different molecular techniques, including looking at organelle variation [10–15]. These previous studies allowed the detection of seven main cpDNA lineages in the O. europaea complex (for the olive tree classification see [16]): lineage E1 was detected in the Mediterranean area and Saharan Mountains, lineages E2 and E3 were specific to the Western Mediterranean area, lineage M was only detected in Macaronesia, lineages C1 and C2 were observed from Southern Asia to Eastern Africa, and lineage A was characteristic of Tropical African olives [15]. One limitation encountered during these studies was the particularly low level of cpDNA and mtDNA polymorphism in the Mediterranean olive tree. Until now only seven haplotypes have been detected with different combinations of loci [17, 18]. These haplotypes belong to lineages E1, E2 and E3 (i.e., two or three haplotypes per lineage [15]). Recently, the first olive plastid genome (cpDNA) was released [18]. For detecting polymorphism in the cultivated olive tree, Mariotti and co-workers analysed sequence variation in 21 cpDNA fragments [18]. Variable microsatellites (also known as simple sequence repeats; SSR), insertions/deletions (indels) in repeated or non-repeated regions, and single nucleotide polymorphisms (SNPs) were identified and allowed for the identification of six cpDNA haplotypes (or chlorotypes) on a set of 30 cultivated olive trees, but they did not find new variants compared to previous studies [17]. The low cpDNA variation detected in the Mediterranean lineages hampered any applications of these markers, particularly for traceability or authenticity of olive oils [17]. Such a low level of cpDNA polymorphism has already been observed for other cultivated woody species such as Prunus avium [19], Vitis vinifera [20] and Pinus pinea [21]. This is probably due to human dispersal of cultivated genotypes originating from a reduced gene pool. In addition, low cpDNA polymorphism has also been reported in forest trees and this may also stem from low mutation rate in long-living organisms [22–24]. However, higher cpDNA variation has been detected in wild olives than in cultivars, and this allowed some population genetic analyses, for instance in the laperrinei and guanchica subspecies from Saharan Mountains and Canary Islands, respectively [25–27].
Additional investigations are needed to maximise the cpDNA haplotype identification in olive trees by testing new markers (especially multiallelic microsatellites [28]) on representatives of both cultivated and wild pools. Here, we address this challenge. Firstly, we sequenced the complete plastid genomes of seven O. europaea accessions, including one Spanish cultivar ('Manzanilla de Sevilla') and six wild olive trees. These taxa were chosen to represent the seven lineages previously reported in the olive tree complex [15]. We also report the complete plastid genome of O. woodiana, a taxon belonging to sect. Ligustroides, which is the sister clade to O. europaea [29]. Secondly, based on these genome sequences, we developed a method for a rapid and routine characterisation of length variation in 62 regions plus two cleaved amplified polymorphism sequence loci (CAPS). A set of 186 cultivars (including both major varieties and local types) as well as five distant wild olive tree populations (129 individuals) were characterised using this approach. Based on the observed polymorphism, we propose an optimised set of primers to detect Mediterranean haplotypes. We also discuss the utility of this approach for forensic analysis as well as for phylogeographic analyses of the olive tree complex.
Results and Discussion
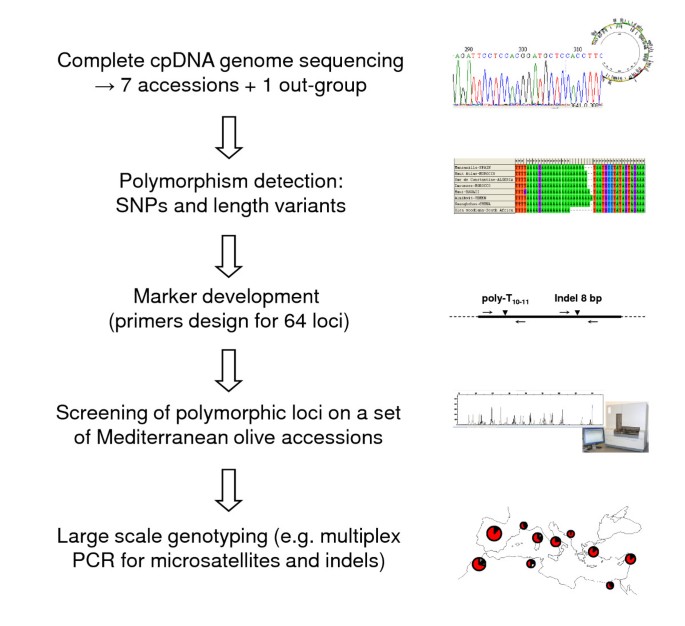
In this study, eight complete olive tree plastid genomes were sequenced and deposited in GenBank/EMBL under the accession numbers FN650747, FN996943, FN996944, FN996972, FN997650, FN997651, FN998900 and FN998901. Polymorphisms were used for the development of new markers to scan cpDNA variation. These loci were used to characterise both cultivated and wild olive trees to assess their utility for forensic and phylogeographic studies. Our general approach is summarised in Figure 1.
Variation in olive tree chloroplast genomes
The cpDNA genome sizes vary between 155,531 base pairs (bp; lineage C2; Almhiwit 5.1) and 155,896 bp (lineage M; Imouzzer S1). As suspected by Besnard & Bervillé [30] based on RFLPs, two long indels were observed in the seven olive tree cpDNA genomes: a 342-bp deletion (in the ycf1 gene) was observed in lineage E3 (Gué de Constantine 20), while a 225-bp deletion (in the trnQ-rps16 intergenic spacer) was detected in both individuals from South Asia (lineages C1 and C2). In addition, 15 smaller indels (i.e., inferior or equal to 12 bp, excluding microsatellite motifs) were also detected. Five of these indels correspond to the presence/absence of a repeated motif of seven to 12 bp (i.e., composed of one or two motifs; located at nucleotide 7,328, 9,526, 14,693, 83,196 and 85,059 in the 'Manzanilla de Sevilla' sequence; see GenBank/EMBL accession no FN996972).
Sequence variation was low, with a total of 218 substitutions on the seven olive plastid genomes. A maximum of 106 substitutions (0.07%) was detected between Gué de Constantine 20 (Algeria) and Almhiwit 5.1 (Yemen), while cpDNA genomes of Guangzhou 1 (China) and Almhiwit 5.1 (Yemen) only showed 34 substitutions (Additional file 1). The plastid genome of O. woodiana displays between 417 and 432 substitutions (< 0.28%) when compared to the seven O. europaea genomes. Again, this level of variation is surprisingly low if we consider that the divergence between sections Olea (O. europaea) and Ligustroides (O. woodiana) is estimated to be between 14 and 22 million years (My; [29]). Based on these results, the cpDNA substitution rate was estimated to be between 1.2 × 10-10 and 2 × 10-10 in the Olea subgenus, which is about ten times lower than the typical mutation rate reported for the plastid genome [7]. This slow molecular evolution might be related to the long generation time of the olive tree [23, 24].
Twelve differences (i.e., three length polymorphisms and nine SNPs, of which one is located in the inverted repeat) were observed between the genomes of 'Frantoio' (GenBank/EMBL accession GU931818; Italy; [18]) and 'Manzanilla de Sevilla' (Spain; this study). According to our approach, we re-sequenced the variable regions in 'Frantoio', from the Olive World Germplasm Bank (OWGB) at Córdoba, Spain (GenBank/EMBL accessions no. FR754486 to FR754495), but these polymorphisms were not confirmed. These 12 differences are not located in the cpDNA regions screened for sequence variation by Mariotti et al. [18] and may be seen as putative sequencing mistakes in accession GU931818. Considering this fact, our analyses indicate that 'Frantoio' and 'Manzanilla de Sevilla' display the same plastid genome, supporting a common maternal origin for these two cultivars.
Based only on nucleotide substitutions (i.e., only 65 out of 218 substitutions were parsimony-informative in the olive tree complex), phylogenetic relationships were depicted from the complete cpDNA genomes using both maximum parsimony (MP) and maximum likelihood (ML) techniques (Figure 2). The resulting topologies confirm results from Besnard et al. [15, 29] through the recovery of two main clades: a Mediterranean/North African clade (clade Cp-II) including lineages E1, E2, E3 and M, and a cuspidata clade (clade Cp-I) including lineages C1, C2 and A. In clade Cp-II, moderate bootstrap support for an early-diverging position of lineage E3 (Gué de Constantine 20) agrees with results based on a few cpDNA microsatellites, indels and CAPS [15]. A moderate level of support was also recovered for the clustering of lineages E1 and E2. Only nine informative substitutions were detected in clade Cp-II, three of them being non-synonymous (Table 1). The information brought by these sites does not strongly support any relationship, suggesting that some sites may be homoplastic. Indeed, two of the three non-synonymous substitutions (52,165 and 83,304) are polymorphic in both clades Cp-I and Cp-II, suggesting that these sites could be under selective pressures, either maintaining polymorphism or contributing to the recurrent appearance of the same substitution (see also [18]). Understanding the molecular variation at these non-synonymous sites would deserve the design of an experiment to test their origin and their adaptive significance.
Plastid DNA phylogenetic tree of the seven olive tree lineages based on nucleotide substitutions from complete plastid genomes. The same topology was obtained with maximum parsimony and maximum likelihood (GTR+I+G) analyses. The bootstrap values are given on each branch (when superior to 50%), the first corresponding to the MP analysis and the second (in brackets) to the ML analysis. The Forsythia europaea and Olea woodiana sequences were used as outgroups. The tree was rooted with the Forsythia sequence. The two clades Cp-I and Cp-II are indicated according to Besnard et al. [15].
Development of cpDNA markers
The low cpDNA substitution rate combined with possible selective effects (which can be problematic for phylogenetic reconstructions [31]) led us to focus on "length polymorphisms". Such polymorphisms were either the result of a variable number of repeats in a microsatellite motif (referred as "microsatellites"), or another type of insertion/deletion (referred as "indel"). Sixty-two regions, of which 51 display variable microsatellite motifs, were investigated (Additional file 2). These sites are located in non-coding regions (except for loci 61 in ycf1) and can thus be considered as mostly neutral. The list of polymerase chain reaction (PCR) primers to amplify the 62 regions is given in Additional file 2. Two CAPS loci (located in rpl14 and the petA-psbJ intergenic spacer) were also characterised to allow the distinction of new haplotypes in lineage E1 (see Methods). After the characterisation of 315 cultivated and wild trees, a multilocus profile (or cpDNA haplotype) was defined for each individual (Additional file 3a). Also, an 88-year old herbarium leaf sample was successfully characterised, suggesting that our method is appropriate for investigating cpDNA variation even on poorly preserved DNA. A total of 40 loci were polymorphic in the Mediterranean/North African olive tree (Additional file 3b). We hope that data generated using this method by different laboratories could be compared to generate a reference dataset for the Mediterranean olive tree. In this way, it should be possible to reconstruct a detailed phylogeography of the species based on a large number of populations, as has been done, for instance, for the European white oaks [32].
Polymorphism assessment in the Mediterranean olive
Some olive tree varieties are used to produce high-quality (and thus more expensive) extra virgin olive oil. Therefore, they may be granted a label of protected designation of origin (PDO; a European Union label referring to food products specific to a particular region or town, conveying a particular quality or characteristic of the specified area). Our markers could find some applications in the traceability of such high quality olive oils, but their discriminating power needs to be determined for assessing their putative utility. Using our cpDNA loci, 12 haplotypes were detected in cultivars (Table 2, Figure 3a and Additional file 3): hence our approach permitted a two-fold increase of the number of detected variants compared to previous studies [17, 18]. The most frequent haplotype (E1.1) was detected in 77% of cultivars, including 'Frantoio' and 'Manzanilla de Sevilla'. Two other haplotypes (E1.2 and E3.2) displayed a frequency superior to 5%, but the remaining haplotypes were rare, and sometimes detected only once (i.e., L1.1, E2.3, E2.5 and E2.6) or twice (i.e., E1.3, E2.2 and E3.1). Several of these rare haplotypes were detected in local cultivars with a limited economic importance (e.g., E2.5, E2.6 and L1.1). The probability that two samples chosen at random display a different haplotype was low (D = 0.40) when compared to nuclear markers, especially nuclear microsatellites for which the discriminating power per locus generally exceeds 0.70 [33–35]. This indicates that the utility of the cpDNA variation for forensic analysis is restricted to rare haplotypes such as the ones detected for 'Picholine' (E2.1) and 'Olivière' (E3.1) in France, 'Villalonga'-'Blanqueta' (E1.3), 'Farga' (E3.1) and 'Lechín de Sevilla' (E2.3) in Spain, or 'Megaritiki' (E2.2) in Greece. These varieties are used to produce high quality extra virgin olive oil (e.g., for Spanish cultivars see [36]). The cpDNA variation, which is a priori easily analysable compared to nuclear single-copy genes, should thus be helpful to complement other procedures for olive traceability based on nuclear polymorphisms [e.g., [37]].
Plastid DNA variation in the Mediterranean olive trees. A. Distribution of the cpDNA haplotypes in cultivated olive trees (see also Additional file 5 for the list of cultivars and the corresponding cpDNA haplotype). B. Distribution of haplotypes in the five studied oleaster populations. For both cultivated and wild gene pools, the number of accessions (n) and the discriminating power (D, D total ) of cpDNA variation is given for each region or population and on the global sample. C. Reduced-median network [54] of cpDNA haplotypes. The traits on branches represent each individual change. Indels are specifically distinguished by bigger orange traits. Each haplotype is represented by a symbol with a definite colour. The name of each cpDNA clade or lineage is given according to Besnard et al. [15] (see also Figure 2). The missing, intermediate nodes are indicated by small black points. CAPS-XapI and CAPS-EcoRI were not considered in this analysis. For this reason, three pairs of haplotypes (i.e., E1-1/E1-4, E1-2/E1-5 and E2-1/E2-4) are not distinguished in the network. In addition, the nine haplotypes not restricted with XapI are indicated with a red circle. * haplotypes for which a complete genome was released in the present study.
In the five populations of oleasters, 18 cpDNA haplotypes were detected, ten of which were shared with cultivars (Table 2, Figure 3b and Additional file 3). The discriminating power of cpDNA was high in these populations (D = 0.89) compared to the cultivated olive tree. Fourteen haplotypes were unique to one population, while the four remaining haplotypes were shared between at least two populations: E1.1 (Rajo, Gialova, Pugnochiuso and Bin El Ouidane), E2.1 and E2.2 (Bin El Ouidane and Pugnochiuso) and E2.3 (Minorca and Bin El Ouidane). These four haplotypes have been detected in cultivated olive trees and could reflect long-distance gene flow mediated by humans [15, 38]. In this way, the most frequent haplotype in cultivars (E1.1) is also the most frequent and widespread haplotype in oleasters (22%; Figure 3b).
Implications for phylogeography
Previous cpDNA phylogeographic studies of the Mediterranean olive tree have been limited due to the low number of haplotypes detected [17, 18]. Here, we demonstrate that a genomic profiling approach of the plastid DNA mostly based on microsatellites and indels can solve this problem. The high variation detected in five distant wild populations indicates a high potential of our approach for resolving the Mediterranean olive tree history. One putative limitation is the level of homoplasy on microsatellite motifs, reported by different authors [39–42], and which could prove problematic when accurately identifying evolutionary relationships between haplotypes. We reconstructed a reduced median network based on molecular markers (Figure 3c). The Mediterranean haplotypes clustered into three lineages (E1, E2 and E3), while the haplotype of subsp. maroccana formed a fourth lineage (M) in northern Africa. This topology is fully congruent with Besnard et al. [15, 29], who used different cpDNA data (i.e., microsatellites, indels and CAPS, or nucleotides). Each lineage displays at least one specific indel, with the exception of lineage M (Figure 3c). Phylogenetic relationships remain unresolved at the base of lineages E1 and E2, as well as in the centre of the network, as a consequence of homoplasy between haplotypes belonging to different lineages (e.g., shared length polymorphisms between clades Cp-I and Cp-II at loci 1, 2, 9, 17, 25, 38, 47, 48, 49, 50 and 58; Additional file 3). Such a difficulty for determining the ancestral state hampers the correct identification of historical links between divergent lineages. In contrast, we expect that homoplasy will not be a serious limitation to resolve phylogenetic relationships among lineages, since their haplotypes have diverged more recently [42]. In any case, for an optimal analysis of the cpDNA variation at the population level, possible length homoplasy will need to be considered and the use of appropriate models will be necessary [41, 43].
The partial or complete cpDNA sequencing of new individuals may reveal nucleotide substitutions that would be of interest [18] for the development of new molecular markers like SNPs (or CAPS). Such SNPs could be used to improve our approach. Nevertheless, the homoplasy is not restricted to repetitive sequences as illustrated with non-synonymous sites in genes under selection, such as the polymorphism detected at the CAPS-XapI locus (in rpl14; Table 1). In the present study, we found restriction polymorphism at this locus in lineages E1 and E2 (clade Cp-II) and also in clade Cp-I (for which we analysed only three accessions; Figure 3c) indicating that this site is highly homoplastic (see also Mariotti et al. [18]). Thus, this site should be used with caution for phylogeographic purposes. Nevertheless, we consider that it could bring potentially important information at the lineage level, particularly to solve the origin of haplotype E1.2 in the cultivated gene pool (7% of cultivars).
Conclusions
A set of 40 polymorphic loci (including 35 with microsatellite motifs) is released for a rapid cpDNA characterization of the Mediterranean olive tree germplasm (see Methods, and Table 3). We expect that, besides their potential forensics application, their use will be important for phylogeographic analyses. Particularly, such studies should allow testing for the persistence of relict populations in the Mediterranean Basin [44], as well as to test the hypotheses about their post-glacial expansion and subsequent domestication [15, 45]. In addition, the identification of genuinely wild populations may represent a significant evolutionary heritage for the conservation of the Mediterranean olive tree diversity. Lastly, the combined use of both nuclear and cpDNA resources should be useful to disentangle the impact of gene dispersal by seeds and pollen on the structure of the genetic diversity. For example, our cpDNA markers will have applications for a comparative study of the dynamic of wild olive tree populations in different environments, such as archipelagos and Saharan mountains [25, 26]. Such information may be relevant for defining appropriate strategies of prospection and in situ conservation of the wild olive tree.
Methods
The general approach is summarised in Figure 1.
Chloroplast genome sequencing
In order to maximize polymorphism detection, the analysis focused on seven individuals of O. europaea L. (subgenus Olea sect. Olea, or olive tree complex), which were chosen to represent one haplotype of each previously described lineage [15]. The following genotypes were thus investigated: 'Manzanilla de Sevilla' (Spanish cultivar; lineage E1), oleaster "Haut Atlas 1" (Morocco; lineage E2), oleaster "Gué de Constantine 20" (Algeria; lineage E3), subsp. maroccana "Imouzzer S1" (Morocco; lineage M), subsp. cuspidata "Maui 1" (Hawaii; lineage A), subsp. cuspidata "Guangzhou CH1" (China; lineage C1), and subsp. cuspidata "Almhiwit C5.1" (Yemen; lineage C2). In addition, we characterised one outgroup species [O. woodiana Knobl. subsp. woodiana (South Africa); sect. Ligustroides Benth. & Hook.], which belongs to the sister group of O. europaea [16, 29]. Appropriate PCR primers were designed to amplify 105 overlapping cpDNA fragments (Additional file 4). Each PCR reaction (25 μl) contained 10 ng DNA template, 1× reaction buffer, 2 mM MgCl2, 0.2 mM dNTPs, 0.2 μmol of each primer, and 0.75 U of Taq DNA polymerase (Promega, Madison, WI, USA). The reaction mixtures were incubated in a thermocycler (T1; Biometra, Göttingen, Germany) for 2 min at 95°C, followed by 36 cycles of 30 s at 95°C (denaturing), 30 s at the annealing temperature (Additional file 4), and 2 min at 72°C (extension). The last cycle was followed by a 10-min extension at 72°C. Direct sequencing of PCR amplicons was performed with an ABI Prism 3100xl Genetic Analyzer, using the Big Dye v3.1 Terminator cycle-sequencing kit, according to the manufacturer's instructions (Applied Biosystems, Foster City, CA, USA). Additionally, nested (internal) primers were also designed to complete the sequencing of each fragment (Additional file 4). The eight Olea genomes were thus reconstructed using a similar approach to the one used by Mariotti et al. [18].
Characterisation of cpDNA polymorphisms in the Mediterranean olive tree
Based on the seven O. europaea sequences, length polymorphism was detected in 62 regions. These polymorphisms were either due to a variable number of repeats in a microsatellite motif or another type of indel (Additional file 2). The PCR primers were designed in flanking regions to specifically amplify short segments (generally inferior to 240 bp). For locus multiplexing, the annealing temperature of all these primers needed to be similar, while the size of PCR products of each locus should be as different as possible. Finally, these primers were also designed to allow amplification of short DNA segments for characterization of poorly preserved material and highly degraded DNAs from herbarium samples. Additionally, the 5' end of the reverse primer of locus 19 was tagged with the sequence GTGTCTT to minimize band stuttering. All primer pairs and specific characteristics of generated fragments are given in Additional file 2. To reduce the cost of the PCR characterization (i.e., time and costs), we used the method described by Schuelke [46]. For each locus (except loci 8, 10, and 61), an 18-bp tail of M13 was added on the forward primer (Additional file 2). When each locus was amplified separately, each PCR reaction (25 μl) contained 10 ng DNA template, 1× reaction buffer, 2.5 mM MgCl2, 0.2 mM dNTPs, 0.2 μmol of one universal fluorescent-labelled M13(-21) primer (5'-TGTAAAACGACGGCCAGT-3'; labelled with one of the three following fluorochromes: HEX, 6-FAM or NED), 0.2 μmol of the reverse primer, 0.05 μmol of the forward primer, and 0.5 U of Taq DNA polymerase (Promega). The reaction mixtures were incubated in a T1 thermocycler for 2 min at 95°C, followed by 28 cycles of 30 s at 95°C, 30 s at 57°C, and 1 min at 72°C, and then by 8 cycles of 30 s at 95°C, 30 s at 51.5°C, and 1 min at 72°C. The last cycle was followed by a 20-min extension at 72°C. Usually, we amplified five or six loci in the same reaction, but in this case, the MgCl2 concentration was increased to 5 mM, and the concentration of primers (except the labelled M13 primer) was decreased by five or six. Loci 8, 10, and 61 (without the M13 tail) were amplified separately with the following conditions: each PCR reaction (25 μl) contained 10 ng DNA template, 1× reaction buffer, 2 mM MgCl2, 0.2 mM dNTPs, 0.2 μmol of each primer, and 0.75 U of Taq DNA polymerase. The reaction mixtures were incubated in a T1 thermocycler for 2 min at 95°C, followed by 36 cycles of 30 s at 95°C, 30 s at 53°C, and 2 min at 72°C. The last cycle was followed by a 10-min extension at 72°C.
The PCR products labelled with a fluorochrome were mixed together with GeneScan-500 ROX as internal standard to run the maximum of loci at the same time (considering the colour and the expected allele size range). They were separated on an ABI Prism 3100xl Genetic Analyzer and the fragment size was determined with GeneMapper version 4.0. For the two non-labelled loci 8 and 61, indels of 342 and 225 bp were revealed under UV after migration on a 2.5% agarose gel electrophoresis stained with GelRed (Biotium, Hayward, CA, USA).
We also focused on the characterisation of two substitutions, which were detected by Mariotti et al. [18] in lineage E1 (the most frequent one in cultivated olive trees; see [13, 17]) and may be potentially useful for forensic analyses and the study of olive tree domestication. We chose to develop two Cleaved Amplified Polymorphism Site (CAPS) loci as in Besnard et al. [47], in order to rapidly characterise a high number of individuals. The PCR primers are given in Additional file 2. The two loci were amplified following the same PCR conditions as for microsatellites. The PCR products were digested with a restriction enzyme (EcoRI or XapI) according to the manufacturer recommendations. The restricted fragments of the two loci were then mixed (with the internal standard ROX 500) and separated on an ABI Prism 3100 xl Genetic Analyzer. The polymorphism for the presence/absence of a restriction site was scored for each genotype. The possibility of multiplexing three different colours (e.g., NED, FAM and HEX) allows the characterisation of 288 (96 × 3) samples per run.
We then characterised 186 cultivated olive tree accessions from different areas with the 64 loci (Table 2, Figure 3a and Additional file 5). The analyzed germplasm includes 106 cultivars from the OWGB Córdoba [48]. These cultivars represent major cultivars from all Mediterranean countries. A few local cultivars from different places were also included in our study for a better representativeness of the cultivated gene pool. First, we characterized 55 cultivated local forms from Morocco (41) and Corsica-Sardinia (14) previously genotyped with nuclear markers [49, 50]. In addition, cultivated trees with or without known denominations from Algeria-Tunisia (6), Italy (6), France (2), Greece-Turkey (3), the Levantine region (5), Libya-Egypt-Sudan (2) and South Africa (1) were added to this study. Beforehand, we tested with nuclear microsatellites that these latter accessions were genetically different (G. Besnard, unpubl. data), except for one herbarium leaf sample from Kufra, Libya (Newberry, sn; 1933 - Kew Herbarium). In addition, to assess the cpDNA variation in the wild Mediterranean olive trees, 129 individuals from five distant populations (Figure 3b) were also characterized: Rajo (Syria; 36°43'50''N, 36°40'00''E), Gialova (Greece; 36°55'12''N, 21°42'42''E), Pugnochiuso (Italy; 41°47'46''N, 16°10'05''E), Minorca (Spain; 39°56'52''N, 04°14'42''E) and Bin El Ouidane (Morocco; 32°03'00''N, 06°35'00''W). To test the reproducibility of the method, the characterisation of ten accessions (i.e., 'Picholine Marocaine', 'Manzanilla de Sevilla', 'Frantoio', 'Moraiolo', 'Ciarasina', 'Confetto', 'Itrana', 'Giaraffa', 'Kalamon' and 'Souri') were repeated three times at random.
Based on this analysis of wild and cultivated accessions, 40 polymorphic loci were detected in the Mediterranean olive trees (Additional file 3). We first proposed to combine 36 of these loci for a rapid characterisation of Mediterranean olive tree germplasm. The multiplex PCRs of five or six loci are proposed in Table 3, but this can be easily modified. The PCR conditions are those previously reported (with the M13 primer). After PCR, these products are mixed together (with no overlap for allele size between loci in a given colour). The locus 10, which needs to be amplified separately, is combined with these multiplex PCRs. Second, when amplified in a multiplex PCR, we encountered some difficulties with locus 19 (not reported in Table 3), and we thus recommend to use it separately and to combine it with the two CAPS (CAPS-XapI and CAPS-EcoRI) for a second combination of three loci. Lastly, the locus 61 is independently characterised on 2.5% agarose gel electrophoresis.
Data analysis
A phylogenetic tree based on the complete plastid genomes was constructed. A partial cpDNA sequence of Forsythia (DQ673256; [51]) was used as an outgroup to root the tree. Sequences were aligned with the application MEGA v4.1 [52]. The alignment was manually refined. Firstly, a maximum parsimony analysis was performed. All characters were equally weighted. The gaps were treated as missing data. A heuristic search was used to find the most parsimonious trees. The close-neighbor-interchange algorithm was used with a search level of 3, as recommended and implemented in the software [52]. The searches included 100 replications of random addition sequences. All the best trees were retained. A strict consensus tree was generated from the equally most-parsimonious trees. The bootstrap values were computed using 10,000 replicates. Secondly, the tree inference was made under a maximum likelihood criterion, using the application PHYML v3.0 [53]. The best-fit substitution model, determined through hierarchical likelihood ratio tests, was the GTR model, with invariable sites and a gamma shape parameter estimated from the data. Support values were obtained by 1,000 bootstrap replicates. Based on fragment genotyping (i.e., microsatellites and indels), the relationships among cpDNA haplotypes were visualized by constructing a reduced median network implemented in the application NETWORK v4.112 [54]. Multi-state microsatellites were treated as ordered alleles and coded by the number of repeated motifs for each allele (e.g., number of T or A; see also [15]) whereas the presence or absence of other indels was coded as 1 and 0, respectively. Basically, this coding strategy assumes that variation at cpDNA microsatellites is mainly due to single-step mutations (e.g., [15, 18]), while allowing consideration of length polymorphisms (microsatellites or indels) with similar weight. However, whether we used different weights or not for indels versus microsatellites did not affect the topology. In addition, for loci combining indels and microsatellite motifs (loci 10, 11, 54 and 57), we separately coded the two types of characters based on available sequences for these loci. The matrix used for the analysis is given in Additional file 6.
The probability that two individuals taken at random display a different haplotype was computed as D = 1 - Σ p i 2, where p i is the frequency of the haplotype i. This parameter was calculated separately on cultivated and wild olive trees, but also on sub-samples or populations. The groups of cultivated olive trees were defined according to their geographic origin.
References
Weir BS, Anderson AD, Hepler AB: Genetic relatedness analysis: modern data and new challenges. Nat Rev Genet. 2006, 7: 771-780. 10.1038/nrg1960.
Lahaye R, Van der Bank M, Bogarin D, Warner J, Pupulin F, Gigot G, Maurin O, Duthoit S, Barraclough TG, Savolainen V: DNA barcoding the floras of biodiversity hotspots. Proc Natl Acad Sci USA. 2008, 105: 2923-2928. 10.1073/pnas.0709936105.
Schaal BA, Olsen KM: Gene genealogies and population variation in plants. Proc Natl Acad Sci USA. 2000, 97: 7024-7029. 10.1073/pnas.97.13.7024.
Petit RJ, Kremer A, Wagner DB: Finite island model for organelle and nuclear genes in plants. Heredity. 1993, 71: 630-641. 10.1038/hdy.1993.188.
Bendich AJ: Why do chloroplasts and mitochondria contain so many copies of their genome?. BioEssays. 1987, 6: 279-282. 10.1002/bies.950060608.
Alaeddini R, Walsh SJ, Abbas A: Forensic implications of genetic analyses from degraded DNA: A review. Forensic Sci Int Genet. 2010, 4: 148-157. 10.1016/j.fsigen.2009.09.007.
Wolfe KH, Li WH, Sharp PM: Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci USA. 1987, 84: 9054-9058. 10.1073/pnas.84.24.9054.
Drouin G, Daoud H, Xia J: Relative rates of synonymous substitutions in the mitochondrial, chloroplast and nuclear genomes of seed plants. Mol Phylogenet Evol. 2008, 49: 827-831. 10.1016/j.ympev.2008.09.009.
Zohary D, Spiegel-Roy P: Beginnings of fruit growing in old world. Science. 1975, 187: 319-327. 10.1126/science.187.4174.319.
Angiolillo A, Mencuccini M, Baldoni L: Olive genetic diversity assessed using amplified fragment length polymorphisms. Theor Appl Genet. 1999, 98: 411-421. 10.1007/s001220051087.
Amane M, Lumaret R, Hany V, Ouazzani N, Debain C, Vivier G, Deguilloux MF: Chloroplast-DNA variation in cultivated and wild olive (Olea europaea L.). Theor Appl Genet. 1999, 99: 133-139. 10.1007/s001220051217.
Besnard G, Khadari B, Villemur P, Bervillé A: Cytoplasmic male sterility in the olive (Olea europaea L.). Theor Appl Genet. 2000, 100: 1018-1024. 10.1007/s001220051383.
Besnard G, Khadari B, Baradat P, Bervillé A: Combination of chloroplast and mitochondrial DNA polymorphisms to study cytoplasm genetic differentiation in the olive complex (Olea europaea L.). Theor Appl Genet. 2002, 105: 139-144. 10.1007/s00122-002-0868-6.
Lumaret R, Ouazzani R, Michaud H, Vivier G, Deguilloux MF, Di Giusto F: Allozyme variation of oleaster populations (wild olive tree) (Olea europaea L.) in the Mediterranean Basin. Heredity. 2004, 92: 334-352.
Besnard G, Rubio de Casas R, Vargas P: Plastid and nuclear DNA polymorphism reveals historical processes of isolation and reticulation in the olive tree complex (Olea europaea). J Biogeogr. 2007, 34: 736-752. 10.1111/j.1365-2699.2006.01653.x.
Green PS: A revision of Olea L. Kew Bull. 2002, 57: 91-140. 10.2307/4110824.
Besnard G: Chloroplast DNA variations in Mediterranean olive. J Hort Sci Biotechnol. 2008, 83: 51-54.
Mariotti R, Cultrera NGM, Muñoz Díez C, Baldoni L, Rubini A: Identification of new polymorphic regions and differentiation of cultivated olives (Olea europaea L.) through plastome sequence comparison. BMC Plant Biol. 2010, 10: 211-10.1186/1471-2229-10-211.
Mohanty A, Martín JP, Aguinagalde I: A population genetic analysis of chloroplast DNA in wild populations of Prunus avium L. in Europe. Heredity. 2001, 87: 421-427. 10.1046/j.1365-2540.2001.00922.x.
Arroyo-García R, Ruiz-García L, Bolling L, Ocete R, López MA, Arnold C, Ergul A, Söylemezoglu G, Uzun HI, Cabello F, Ibáñez J, Aradhya MK, Atanassov A, Atanassov I, Balint S, Cenis JL, Constantini L, Goris-Lavets S, Grando MS, Klein I, McGovern PE, Merdinoglu D, Pejic I, Pelsy F, Primkirios N, Risovannaya V, Snoussi A, Sotiri P, Tamhankar S, This P, Troshin L, Malpica JM, Lefort F, Martinez-Zapater JM: Multiple origins of cultivated grapevine (Vitis vinifera L. ssp. sativa) based on chloroplast DNA polymorphisms. Mol Ecol. 2006, 15: 3707-3714. 10.1111/j.1365-294X.2006.03049.x.
Vendramin GG, Fady B, González-Martínez SC, Sheng Hu F, Scotti I, Sebastiani F, Soto A, Petit RJ: Genetically depauperate but widespread: The case of an emblematic Mediterranean pine. Evolution. 2008, 62: 680-688. 10.1111/j.1558-5646.2007.00294.x.
Heuertz M, Carnevale S, Fineschi S, Sebastiani F, Hausman JF, Paule L, Vendramin GG: Chloroplast DNA phylogeography of European ashes, Fraxinus sp (Oleaceae): roles of hybridization and life history traits. Mol Ecol. 2006, 15: 2131-2140. 10.1111/j.1365-294X.2006.02897.x.
Magri D, Fineschi S, Bellarosa R, Buonamici A, Sebastiani F, Schirone B, Simeone MC, Vendramin GG: The distribution of Quercus suber chloroplast haplotypes matches the palaeogeographical history of the western Mediterranean. Mol Ecol. 2007, 16: 5259-5266. 10.1111/j.1365-294X.2007.03587.x.
Smith SA, Donoghue MJ: Rates of molecular evolution are linked to life history in flowering plants. Science. 2008, 322: 86-89. 10.1126/science.1163197.
Besnard G, Christin PA, Baali-Cherif D, Bouguedoura N, Anthelme F: Spatial genetic structure in the Laperrine's olive (Olea europaea subsp. laperrinei), a long-living tree from central-Saharan mountains. Heredity. 2007, 99: 649-657. 10.1038/sj.hdy.6801051.
García-Verdugo C, Forrest AD, Ballaguer L, Fay MF, Vargas P: Parallel evolution of insular Olea europaea subspecies based on geographical structuring of plastid DNA variation and phenotypic similarity in leaf traits. Bot J Linn Soc. 2010, 162: 54-63. 10.1111/j.1095-8339.2009.01013.x.
García-Verdugo C, Forrest AD, Fay MF, Vargas P: The relevance of gene flow in metapopulation dynamics of an oceanic island endemic, Olea europaea subsp. guanchica. Evolution. 2010, 64: 3525-3536. 10.1111/j.1558-5646.2010.01091.x.
Ebert D, Peakall R: Chloroplast simple sequence repeats (cpSSRs): technical resources and recommendations for expanding cpSSR discovery and applications to a wide array of plant species. Mol Ecol Resourc. 2009, 9: 673-690. 10.1111/j.1755-0998.2008.02319.x.
Besnard G, Rubio de Casas R, Christin PA, Vargas P: Phylogenetics of Olea (Oleaceae) based on plastid and nuclear ribosomal DNA sequences: tertiary climatic shifts and lineage differentiation times. Ann Bot. 2009, 104: 143-160. 10.1093/aob/mcp105.
Besnard G, Bervillé A: On chloroplast DNA variations in the olive (Olea europaea L.) complex: comparison of RFLP and PCR polymorphisms. Theor Appl Genet. 2002, 104: 1157-1163. 10.1007/s00122-001-0834-8.
Christin PA, Weinreich DM, Besnard G: Causes and evolutionary significance of genetic convergence. Trends Genet. 2010, 26: 400-405. 10.1016/j.tig.2010.06.005.
Petit RJ, Brewer S, Bordacs S, Burg K, Cheddadi R, Coart E, Cottrell J, Csaikl UM, van Dam B, Deans JD, Espinel S, Fineschi S, Finkeldey R, Glaz I, Goicoechea PG, Jensen JS, Konig AO, Lowe AJ, Madsen SF, Matyas G, Munro RC, Popescu F, Slade D, Tabbener H, de Vries SGM, Ziegenhagen B, de Beaulieu JL, Kremer A: Identification of refugia and post-glacial colonisation routes of European white oaks based on chloroplast DNA and fossil pollen evidence. Forest Ecol Manage. 2002, 156: 49-74. 10.1016/S0378-1127(01)00634-X.
Besnard G, Breton C, Baradat P, Khadari B, Bervillé A: Cultivar identification in olive based on RAPD markers. J Amer Soc Hort Sci. 2001, 126: 668-675.
Belaj A, Satovic Z, Cipriani G, Baldoni L, Testolin R, Rallo L, Trujillo I: Comparative study of the discriminating capacity of RAPD, AFLP and SSR markers and of their effectiveness in establishing genetic relationships in olive. Theor Appl Genet. 2003, 107: 736-744. 10.1007/s00122-003-1301-5.
Khadari B, Breton C, Moutier N, Roger JP, Besnard G, Bervillé A, Dosba F: The use of molecular markers for germplasm management in a French olive collection. Theor Appl Genet. 2003, 106: 521-529.
ICEX Spain: Olive Oil from Spain. (accessed on 01/12/2010), [http://www.oliveoilfromspain.com/OOFS/everything/denominations_of_origins.asp]
Consolandi C, Palmieri L, Severgnini M, Maestri E, Marmiroli N, Agrimonti C, Baldoni L, Donini P, de Bellis G, Castiglioni B: A procedure for olive oil traceability and authenticity: DNA extraction, multiplex PCR and LDR-universal array analysis. Eur Food Res Technol. 2008, 227: 1429-1438. 10.1007/s00217-008-0863-5.
Besnard G, Khadari B, Baradat P, Bervillé A: Olea europaea (Oleaceae) phylogeography based on chloroplast DNA polymorphism. Theor Appl Genet. 2002, 104: 1353-1361. 10.1007/s00122-001-0832-x.
Doyle JJ, Morgante M, Tingey SV, Powell W: Size homoplasy in chloroplast microsatellites of wild perennial relatives of soybean (Glycine subgenus Glycine). Mol Biol Evol. 1998, 15: 215-218.
Provan J, Powell W, Hollingsworth PM: Chloroplast microsatellites: new tools for studies in plant ecology and evolution. Trends Ecol Evol. 2001, 16: 142-147. 10.1016/S0169-5347(00)02097-8.
Navascués M, Emerson BC: Chloroplast microsatellites: measures of genetic diversity and the effect of homoplasy. Mol Ecol. 2005, 14: 1333-1341. 10.1111/j.1365-294X.2005.02504.x.
Vachon N, Freeland JR: Phylogeographic inferences from chloroplast DNA: quantifying the effects of mutations in repetitive and non-repetitive sequences. Mol Ecol Resourc. 2011, 11: 279-285. 10.1111/j.1755-0998.2010.02921.x.
Estoup A, Jarne P, Cornuet JM: Homoplasy and mutational model at microsatellite loci and their consequences for populations genetics analysis. Mol Ecol. 2002, 11: 1591-1604. 10.1046/j.1365-294X.2002.01576.x.
Carrion Y, Ntinou M, Badal E: Olea europaea L. in the North Mediterranean Basin during the Pleniglacial and the Early-Middle Holocene. Quat Sci Rev. 2010, 29: 952-968. 10.1016/j.quascirev.2009.12.015.
Breton C, Terral JF, Pinatel C, Médail F, Bonhomme F, Bervillé A: The origins of the domestication of the olive tree. C R Biol. 2009, 332: 1059-1064. 10.1016/j.crvi.2009.08.001.
Schuelke M: An economic method for the fluorescent labelling of PCR fragments. Nat Biotechnol. 2000, 18: 233-234. 10.1038/72708.
Besnard G, Rubio de Casas R, Vargas P: A set of primers for length and nucleotide-substitution polymorphism in chloroplastic DNA of Olea europaea L. (Oleaceae). Mol Ecol Notes. 2003, 3: 651-653. 10.1046/j.1471-8286.2003.00547.x.
Caballero JM, del Río C, Barranco D, Trujillo I: The olive world germplasm bank of Córdoba, Spain. Olea. 2006, 25: 14-19.
de Caraffa VB, Maury J, Gambotti C, Breton C, Bervillé A, Giannettini J: Mitochondrial DNA variation and RAPD mark oleasters, olive and feral olive from Western and Eastern Mediterranean. Theor Appl Genet. 2002, 104: 1209-1216. 10.1007/s00122-002-0883-7.
Khadari B, Charafi S, Moukhli A, Ater M: Substantial genetic diversity in cultivated Moroccan olive despite a single major cultivar: a paradoxical situation evidenced by the use of SSR loci. Tree Genet Genomes. 2008, 4: 213-221. 10.1007/s11295-007-0102-4.
Lee HL, Jansen RK, Chumley TW, Kim KJ: Gene relocations within chloroplast genomes of Jasminum and Menodora (Oleaceae) are due to multiple, overlapping inversions. Mol Biol Evol. 2007, 24: 1161-1180. 10.1093/molbev/msm036.
Tamura K, Dudley J, Nei M, Kumar S: Mega4: Molecular evolutionary genetics analysis (Mega) software version 4.0. Mol Biol Evol. 2007, 24: 1596-1599. 10.1093/molbev/msm092.
Guindon S, Gascuel O: A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003, 52: 696-704. 10.1080/10635150390235520.
Bandelt HJ, Forster P, Röhl A: Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol. 1999, 16: 37-48.
Acknowledgements
We thank Virginie Brunini, Christos Mammides, Andriana Minou, Giorgos Minos, Alex Papadopoulos and Carmen del Río (OWGB, IFAPA, Centro Alameda del Obispo, Córdoba, Spain; FEDER-INIA RFP2009-00008-C2-01), who provided olive tree samples or DNA extracts. One leaf sample was also kindly provided by the Kew herbarium. This work was funded by the Intra-European fellowship PIEF-GA-2008-220813 to GB. PH was supported by MICINN grant AGL2010-17316 from the Spanish Ministry of Science and Innovation. GD was supported by projects 041/C/2007, 75/C/2009 & 56/C/2010 of "Consejería de Agricultura y Pesca, Junta de Andalucía"; "Grupo PAI" AGR-248 of "Junta de Andalucía"; and "Ayuda a Grupos" of "Universidad de Córdoba" (Spain). VS was supported by grants from the ERC, Leverhulme Trust, NERC and the Royal Society. We also thank Silvana del Vecchio for lab assistance, Martyn Powell and two anonymous reviewers for helpful comments on this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
GB & VS designed the initial project, with subsequent contributions by the other authors. GB conducted the experiments and wrote the initial version of the manuscript. GD and PH contributed to olive cpDNA sequencing and to the acquisition of cultivated olive genotyping data. BK contributed to the acquisition of data for cultivated and wild olive trees. GB, PH, BK, GD and VS revised the manuscript critically. All authors have given final approval for this version to be published.
Electronic supplementary material
12870_2011_869_MOESM2_ESM.XLS
Additional file 2: Loci features. Primers, allele size range, polymorphism type, genome location and corresponding names in previous studies are given (XLS 58 KB)
12870_2011_869_MOESM3_ESM.XLS
Additional file 3: Plastid DNA variation based on the 64 loci. a) Profiles for the 321 trees characterized in this study (including those for complete cpDNA genomes); and b) Different cpDNA haplotypes. (XLS 190 KB)
12870_2011_869_MOESM4_ESM.DOC
Additional file 4: PCR amplification and sequencing primers (5'->3') used to amplify and sequence the complete olive plastid genome. (DOC 202 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Besnard, G., Hernández, P., Khadari, B. et al. Genomic profiling of plastid DNA variation in the Mediterranean olive tree. BMC Plant Biol 11, 80 (2011). https://doi.org/10.1186/1471-2229-11-80
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2229-11-80